Research
The Principles Governing Embryonic Cell Fate Decisions
At the heart of every living organism's development lies a remarkable journey that begins with a single fertilized egg. During the early stages of embryogenesis, this seemingly uniform cell undergoes a series of intricate and orchestrated decisions that ultimately determine its destiny. These decisions dictate whether a cell will become a nerve cell, a muscle cell, or a myriad of other specialized cell types, setting the foundation for the complex, multicellular life that follows. Understanding the mechanisms behind these early cell fate decisions is not only a fundamental question in developmental biology but also holds profound implications for reproductive medicine, disease modeling, and our comprehension of life itself.
TE vs. ICM: Defining Developmental Paths. In the early stages of embryogenesis, a pivotal decision unfolds as the developing embryo transitions from a single fertilized cell into a blastocyst. Here, cells encounter a decisive crossroads in their developmental trajectory. Some commit to becoming the Trophectoderm (TE), destined for the formation of extraembryonic tissues vital for implantation and placental development. In contrast, others choose the path of the Inner Cell Mass (ICM), which eventually gives rise to the epiblast, shaping the organism's future body, and the Primitive Endoderm (PE), contributing to essential structures like the yolk sac. This initial cell fate decision establishes the foundation for subsequent processes of differentiation and specialization, paving the way for a multitude of intriguing questions:
-
How do cells within the TE and ICM acquire their distinct identities, and what molecular cues govern this decision?
-
What orchestrates the delicate balance between self-renewal and differentiation within the ICM, leading to the emergence of epiblast and PE lineages?
-
Are there critical checkpoints or master regulators that control the fate of these embryonic cells as they embark on their respective developmental journeys?
-
What implications do these early cell fate decisions hold for our comprehension of the origins of life and the intricate processes that drive embryonic development?
These questions serve as gateways to the exploration of an awe-inspiring realm where cells make choices that shape the destiny of living organisms, offering fertile ground for ongoing scientific inquiry and discovery.
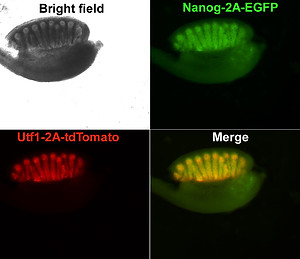
Sex Determination in Bipotential Gonads: Blueprinting Identity. The journey of sexual identity unfolds in parallel with the earliest stages of embryonic development, ushering in a profound exploration of identity blueprinting within bipotential gonads. As these nascent structures take form, poised to evolve into either ovaries or testes, intricate genetic and hormonal processes come into play, decisively determining the embryo's sex. The orchestration of this symphony involves a complex interplay of genes, with pivotal players like SRY in males and FOXL2 in females acting as conductors guiding the gonads toward their distinct destinies.
However, the ramifications of this process extend well beyond the realm of anatomical distinctions. It serves as the catalyst for an intricate cascade of sexual development, shaping secondary sexual characteristics, regulating reproductive function, and even exerting an influence over overall health. This enigmatic journey provokes a series of thought-provoking questions:
-
How does the spatiotemporal expression of SRY is regulated? and which are the most potent SRY activators?
-
Are there windows of plasticity or critical checkpoints within the process of SRY activation where interventions could potentially influence or alter the outcome?
-
How might our expanding understanding of sex determination hold implications for addressing disorders of sexual development and promoting reproductive health?
As we delve into this captivating realm of sexual identity determination, we unearth a treasure trove of questions that not only challenge our understanding of development but also invite us to explore the profound impact of these early decisions on the course of life itself.


The Molecular Basis of Cellular Plasticity
Regenerative medicine stands as a burgeoning frontier, striving to mend the human body by replacing lost or damaged cells, tissues, or organs through the art of cellular transplantation. Within this landscape, embryonic stem cells (ESCs) emerge as remarkable pluripotent entities, possessing the capacity for long-term growth, self-renewal, and the extraordinary potential to give rise to every cell, tissue, and organ in the developing fetus.
At the center of ESCs lies an epigenetic landscape characterized by a "hyperdynamic" chromatin state. This unique feature enables these cells to swiftly and efficiently respond to both external and internal cues. Such cues trigger the activation of key master regulators, driving ESCs toward their respective developmental fates. Consequently, ESCs hold immense promise as a wellspring of diverse differentiated cell types, offering tantalizing prospects for cell therapy.
However, realizing this potential faces significant bottlenecks. Allogenic immune rejection of ESC-derived cells by recipients looms as a formidable challenge, alongside ethical considerations.
Nuclear Reprogramming and Direct Lineage Conversion: Methods of Producing Medically-Relevant Cell Types for the Clinic. The paradigm of cellular plasticity underwent a transformative shift, by demonstrating that the introduction of just four transcription factors, OCT4, SOX2, KLF4, and MYC, collectively known as OSKM, could reprogram adult fibroblasts into functional embryonic stem-like cells, referred to as induced pluripotent stem cells (iPSCs).
While somatic cell nuclear transfer (SCNT) had previously achieved somatic cell epigenome resetting, it necessitated the activation of hundreds to thousands of genes expressed within the oocyte. The revelation that a mere quartet of factors could reset a cell's epigenome opened a novel avenue. Scientists ventured to convert different adult cells into somatic counterparts from divergent ontogenic lineages, bypassing pluripotency. This innovative approach hinged on specific subsets of master regulators.
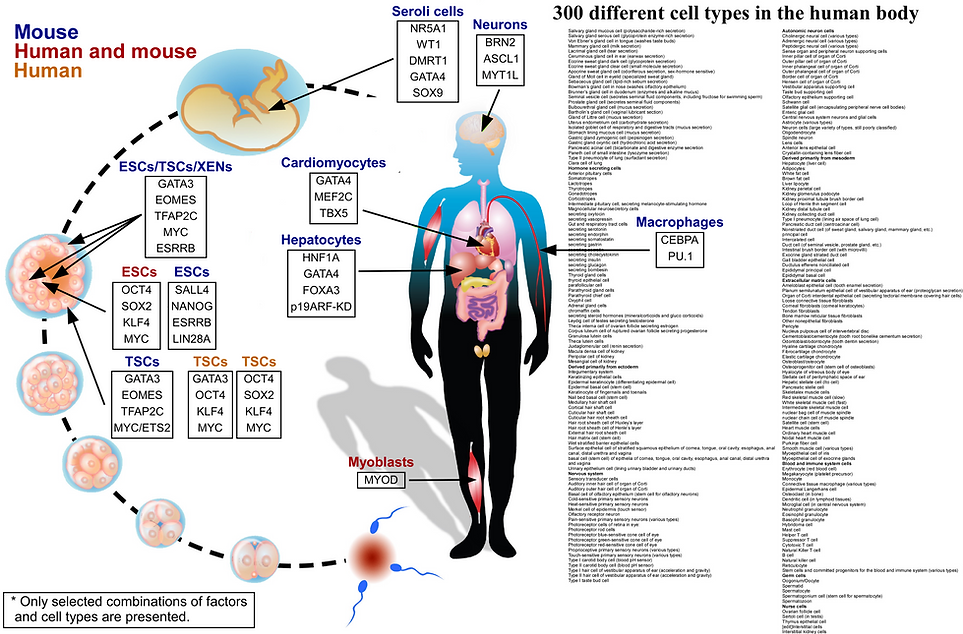
The pursuit of direct conversion led to breakthroughs across a multitude of cell types, including hematopoietic cells, various neuronal subtypes, cardiomyocytes, hepatocytes, embryonic Sertoli cells, endothelial cells, retinal pigment epithelium (RPE), trophoblast stem cells (TSCs), and extraembryonic endoderm stem cells (XENs). Notably, the generation of iPSCs and directly converted cells addressed the challenges of immune rejection and ethical concerns. Yet, challenges persist, as many directly converted cells exhibit instability, representing incomplete reprogramming states, and most iPSCs, while activating their endogenous circuitry, display limited developmental potential in murine models.
These findings underscore the imperative need for refining and enhancing existing reprogramming methodologies before contemplating their application in clinical settings. As we navigate this intricate landscape of cellular reprogramming, we confront a host of questions:
-
How can we achieve stable and complete reprogramming in directly converted cells, unlocking their therapeutic potential?
-
What molecular insights can we glean from the elusive balance between somatic and pluripotent states?
-
Are there undiscovered master regulators that hold the key to successful cellular reprogramming?
These questions propel us further into the captivating realm of reprogramming, where the pursuit of answers promises to reshape the trajectory of regenerative medicine and redefine the boundaries of cellular plasticity.
Partial Reprogramming by Defined Factors: A Cutting-Edge Technology Poised to Revolutionize Cellular Rejuvenation. In recent years, researchers have made remarkable strides in unraveling the potential of partial reprogramming for rejuvenation purposes. In this technique, aged cells are briefly exposed to pluripotency reprogramming factors, allowing them to regain a younger epigenetic landscape and improved function. This technology offers a glimpse into a future where age-related diseases may be mitigated, and damaged tissues repaired.
Yet, within this burgeoning field, numerous questions await answers:
-
Is partial reprogramming to pluripotency the sole method for inducing cellular rejuvenation?
-
Can rejuvenated cells maintain their beneficial properties for an extended period after the removal of the transgenes?
-
What are the full implications of partial reprogramming for the long-term health of rejuvenated cells?
-
How can we fine-tune the reprogramming process to ensure safety and efficacy in clinical applications?
-
Can we extend its benefits to a broader range of cell types and tissues?
We embark on a journey through the principles, methodologies, mechanism of action, and milestones achieved in the realm of partial reprogramming by defined factors. The exploration of partial reprogramming continues to illuminate our understanding of cellular dynamics and its transformative potential in the domain of regenerative medicine.

The Fundamental Features of Stemness
Stemness, the defining trait of stem cells, encapsulates the remarkable ability to adapt, differentiate, and regenerate, a fundamental attribute at the heart of development and regenerative medicine. It is the finely tuned interplay of genes and molecules that guides stem cells along their developmental path, enabling them to maintain their undifferentiated state or embark on specialized lineages. The intricate interplay of non-coding RNAs (ncRNAs), X chromosome reactivation/inactivation dynamics, transcription factor modulations, metabolic shift, and epigenetic reprogramming orchestrate the regulatory mechanisms that govern the pluripotent and multipotent stemness states.
The Silent Architects: The Role of lncRNAs in Stemness. Long non-coding RNAs (LncRNAs), traditionally overshadowed by their protein-coding counterparts, have recently taken center stage for their pivotal roles in regulating stemness. These non-coding transcripts exhibit a multifaceted influence over chromatin remodeling, gene expression, and the intricate balance between self-renewal and differentiation. They serve as molecular architects, sculpting the regulatory landscapes that define stem cell identity.
While recent discoveries shed some light on the role of lncRNAs in maintaining stemness, many open questions remained to be answered.
-
How do lncRNAs exert their influence over key cellular processes crucial for stemness?
-
What molecular interactions underlie their ability to fine-tune gene expression and chromatin architecture?
-
Are there stem cell-specific lncRNAs that govern distinct aspects of stemness?
-
How do different lncRNAs contribute to the heterogeneity observed among stem cell populations?
-
Can we harness the regulatory power of lncRNAs for therapeutic interventions in regenerative medicine?
As we navigate this scientific terrain, these unanswered questions beckon us to delve deeper into the molecular intricacies of lncRNAs, offering the potential to reshape our understanding of stemness.
X Chromosome Reactivation: Why Only in Pluripotency? Unlike most female somatic cells, which maintain one inactive X chromosome, pluripotent stem cells employ mechanisms to reactivate and balance X-linked gene expression. The process of X chromosome reactivation, while still enigmatic in many aspects, represents a pivotal axis in female stem cell biology.
This scientific inquiry embarks on an exploration of the role of X chromosome reactivation in stemness, delving into the molecular underpinnings of this intricate phenomenon. Alongside these discoveries, a series of unanswered questions emerges:
-
What are the molecular mechanisms that enable X chromosome reactivation in female pluripotent stem cells?
-
Can these mechanisms be harnessed in other cell types to reactivate the X chromosome?
-
How do these mechanisms differ from those observed in other multipotent stem cells?
-
What functional consequences does X chromosome reactivation have on stem cell identity, differentiation potential, and therapeutic utility?
-
Can insights into X chromosome reactivation be leveraged to enhance regenerative therapies and personalized medicine approaches?
As we navigate the intriguing landscape of X chromosome reactivation in female stemness, these unanswered questions drive us to explore further, unveiling the mysteries that underlie the dynamics of X chromosome reactivation and inactivation.






